RNA interference, RNAi, offers a powerful means of silencing disease-relevant genes by directing short double-stranded RNA, dsRNA, molecules to degrade or stall the translation of target messenger RNA. Longer dsRNA constructs could in principle silence multiple genes simultaneously, but their clinical utility is curtailed by two persistent problems: poor stability in biological environments and inefficient cellular uptake. Peptides derived from natural RNA-binding proteins have emerged as a promising class of dsRNA stabilizers, but how these peptides physically alter the mechanical properties of dsRNA, and what those changes mean for delivery and release, had not been characterized at the single-molecule level.
Researchers in the Dulin Group at Vrije Universiteit Amsterdam investigated the biophysical effects of two peptides derived from the tomato aspermy virus 2b, TAV2b, protein on long dsRNA molecules, published in the Journal of the American Chemical Society. TAV2b naturally suppresses host gene silencing by binding the major groove of dsRNA as a homodimer. The team studied a 33-residue wild-type peptide, wt33, representing the core RNA interaction motif, alongside a homodimeric derivative, 1'-1', in which the native leucine zipper dimerization motif is replaced by a disulfide bond. This redesign introduces a redox-sensitive release mechanism: in the reducing environment of the cytosol, the disulfide bond is cleaved, monomerizing the peptide and releasing the bound dsRNA. Using high-throughput magnetic tweezers, MT, the team applied precisely controlled force and torque to individual dsRNA tethers approximately 3,200 base pairs in length, monitoring changes in contour length, persistence length, stretch modulus, and helical twist in real time before and after peptide addition.
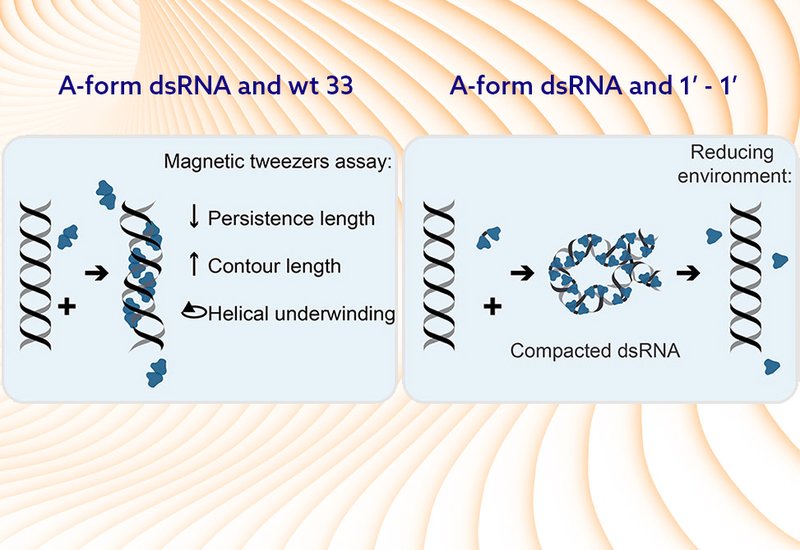
Both peptides underwound dsRNA upon binding, shifting the zero-twist peak position by up to 15 turns at saturation, consistent with structural analysis of the TAV2b-dsRNA crystal structure showing that protein binding widens the major groove from 11 Å to 19 Å and increases the helical periodicity from 11 to 12 base pairs per turn. Peptide wt33 increased the contour length of dsRNA by approximately 10%, reduced the persistence length from 45 nm to 10 nm, and lowered the stretch modulus, collectively indicating that the peptide renders dsRNA more flexible and extended. Real-time binding experiments demonstrated that wt33 associates with and dissociates from dsRNA rapidly and reversibly, consistent with a two-step equilibrium model in which dsRNA fluctuates between double-stranded and locally melted conformations, with wt33 selectively stabilizing the double-stranded form at lower twist. The half-saturation concentration extracted from this model was 1.2 μM. Peptide 1'-1' produced a qualitatively different outcome: at forces below 1 pN, it condensed the dsRNA tether through higher-order peptide-peptide interactions, forming a compacted complex that did not dissociate under buffer washing but released completely upon addition of the reducing agent tris(2-carboxyethyl)phosphine, TCEP.
These findings establish that peptide design directly determines whether a dsRNA binder produces a flexible, dynamically exchanging complex or a condensed, stably packaged one, both of which carry distinct implications for RNA therapeutics. The reduced bending rigidity conferred by wt33 could facilitate the packaging of long dsRNA constructs, including self-amplifying RNA vectors that may reach 10 kilobase pairs in length, into lipid or polymer nanocarriers. The compaction and redox-triggered release behavior of 1'-1' suggests a complementary strategy for protecting dsRNA in circulation and achieving spatiotemporally controlled intracellular delivery. Together, the single-molecule magnetic tweezers framework demonstrated here provides a quantitative toolkit for evaluating and guiding the rational design of next-generation dsRNA binders for gene silencing and antiviral applications.