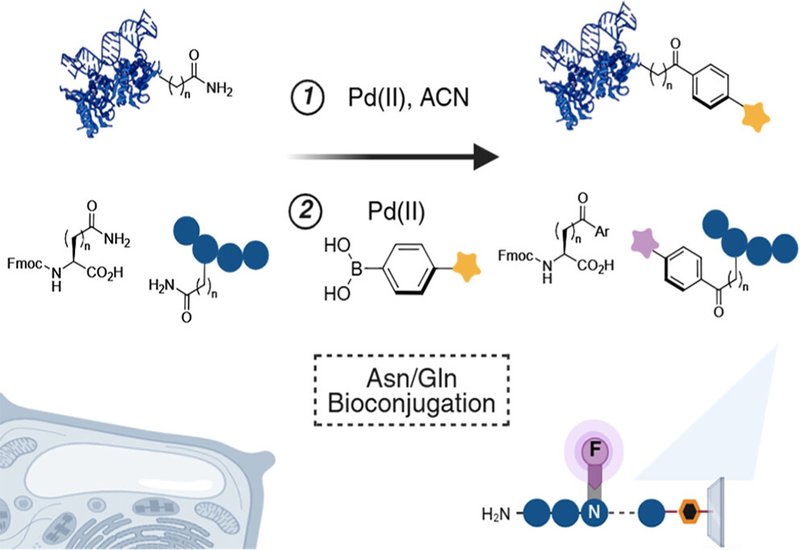
Asparagine, Asn, and glutamine, Gln, together account for roughly 8% of all residues in the human proteome, a proportion comparable to serine, the most abundant amino acid. Despite that prevalence, both residues have remained chemically inaccessible to selective modification. Their neutral primary amide side chains lack the nucleophilicity of thiols, the redox sensitivity of methionine, and the electrophilic handles that established bioconjugation platforms exploit. A further obstacle has confined earlier palladium-mediated dehydration attempts to small, protected substrates: the abundance of heteroatom-rich side chains in native proteins, particularly histidine, lysine, cysteine, and methionine, together with free N-termini, rapidly coordinates Pd(II) into stable, nonreactive chelates that poison the catalyst before any useful chemistry can occur. The result is a chemically silent slice of the proteome that has resisted every general modification strategy.
Researchers in the Raj Group at Emory University, published in the Journal of the American Chemical Society, solved the chelation problem by engineering a reaction sequence that couples kinetically controlled isohypsic dehydration with effective metal scavenging. Using Pd(O2CCF3)2 in a water–acetonitrile mixture, paired with thiol-based scavengers such as L-cysteine or 3-mercaptopropionic acid, 3-MPA, the team neutralized competing chelation by sequestering residual palladium into water-soluble thiolate complexes removable by filtration or dialysis. Residual Pd levels after scavenging fell to 43.9 ppm per milligram of protein, satisfying ICH Q3D pharmaceutical standards and permitting downstream analytical and therapeutic applications. Chemoselectivity studies using a peptide bearing lysine, tyrosine, tryptophan, cysteine, serine, methionine, glutamate, histidine, and arginine confirmed that dehydration and subsequent carbometalation proceed without touching any residue other than Asn and Gln, with greater than 99% conversion achieved using Pd(O2CCF3)2. The resulting nitrile handle, compact and stable, undergoes further Pd-mediated carbometalation with aryl boronic acids to yield aryl ketone products across a broad scope: electron-donating boronic acids couple smoothly at 40–60 °C in 73–89% isolated yield, while electron-withdrawing partners require moderately elevated temperatures.
The platform's reach proved wide. Applied to eight intact proteins spanning 8.5–66.5 kDa, from ubiquitin to bovine serum albumin, BSA, the dehydration step achieved 84–95% Asn and Gln conversion, and circular dichroism spectroscopy confirmed that secondary structure was largely preserved throughout. Subsequent carbometalation installed 2–9 aryl ketone sites per protein, and dual functionalization of BSA with boronic acids bearing azide, alkyne, ketone, or vinyl groups demonstrated the handle's versatility as a platform for attaching diverse affinity and reporter motifs. Applied to the cytotoxic peptide AERQ on solid support, a 4-diphenylamino aryl ketone analogue produced a statistically significant increase in HeLa cell death relative to the parent peptide, illustrating how a single Asn and Gln to ketone conversion generates new pharmacological space without de novo sequence redesign. For single-molecule protein sequencing, Asn residues in model peptides were labeled with azide-ketone handles, coupled to Atto643 fluorophore via copper-free click chemistry, and detected position-specifically by total internal reflection fluorescence microscopy during sequential Edman degradation cycles. The translational reach extended to trastuzumab, a monoclonal antibody targeting the human epidermal growth factor receptor 2, HER2: site-specific modification at N437, Q441, and Q648 followed by carbometalation and hydroxylamine-AlexaFluor647 conjugation produced a labeled antibody that retained strong, specific binding to HER2-overexpressing BT-474 breast cancer cells while generating negligible signal on HER2-negative MCF 10A cells.
By transforming Asn and Gln from passive spectators into programmable entry points for diversification, this platform complements established cysteine, lysine, and methionine bioconjugation chemistries with the first general strategy to exploit neutral primary amide side chains in native biomolecules. A companion study from the Raj Group, published in ACS Chemical Biology, extended this chemistry to proteome-wide scale, deploying the palladium-mediated dehydration platform across complex cell lysates and living cells to map Asn and Gln sites globally and to develop an inverse chemoproteomic framework for profiling deamidation and N-glycosylation post-translational modifications.