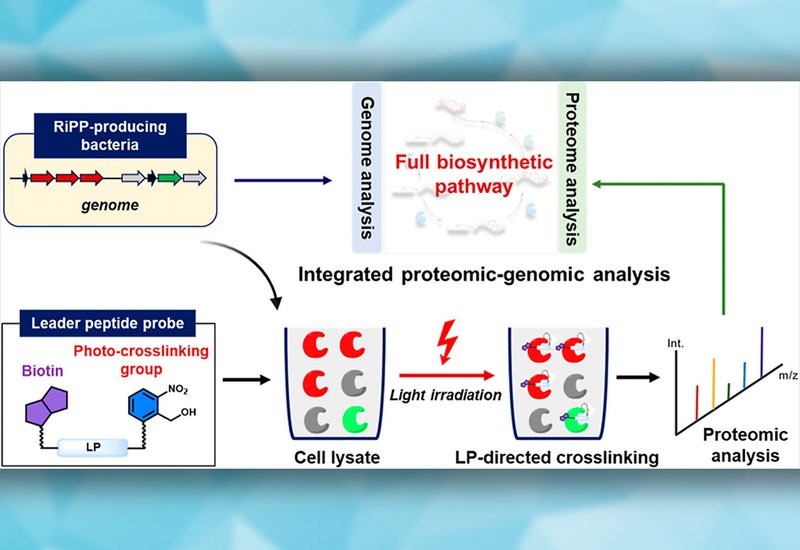
Ribosomally synthesized and post-translationally modified peptides, RiPPs, are a structurally diverse family of natural products whose biosynthesis depends on a conserved logic: an N-terminal leader peptide recruits and activates modification enzymes that act on the adjacent core peptide. Genome mining has transformed RiPP discovery by identifying precursor peptides and tailoring enzymes within co-localized biosynthetic gene clusters, BGCs. Yet the cluster-centric assumption increasingly fails to capture the full enzymatic complement of RiPP pathways. Orphan proteases, promiscuous synthetases, and other auxiliary factors encoded far from canonical BGCs go undetected by sequence-context analysis alone, leaving biosynthetic networks incompletely mapped.
Researchers in the Wang Lab at Nanjing University, published in J. Am. Chem. Soc., addressed this gap by designing covalent leader peptide probes that capture LP-binding enzymes directly from native proteomes. The team attached an o-nitrobenzyl azide photo-cross-linker to the N- or C-terminus of leader peptides from two well-characterized RiPP systems: LctA from lacticin 481 biosynthesis and PatE from the cyanobactin pathway. Upon 365 nm irradiation, the probes formed stable covalent conjugates with their cognate enzymes, LctM and LynD respectively. Microscale thermophoresis confirmed that probe attachment did not perturb leader peptide recognition, and a scrambled-sequence control produced no detectable labeling, establishing sequence-dependent specificity. The team then extended the approach to the RiPP-producing strain Streptomyces sparsogenes ATCC 25498, deploying SpaA1.2-derived probes carrying either an NBD fluorophore or a biotin handle for affinity enrichment followed by LC-MS/MS identification.
Validation with purified proteins confirmed that photo-cross-linking occurs at the authentic LP-binding site: LctM covalently tethered to LctALP could no longer bind free LctALP as measured by microscale thermophoresis, yet the conjugate modified the LctA core peptide with efficiency comparable to the LctCE fusion enzyme, generating fully dehydrated and cyclized products bearing the characteristic lacticin 481 ring topology. The LynD-PatELP conjugate similarly restored robust macrocyclization activity toward the PatE core peptide, a reaction the unconjugated enzyme performs only minimally. These paired results establish that covalent capture preserves, and in the LP-activated case effectively locks in, the catalytically competent state.
Applied to S. sparsogenes lysates, the biotinylated SpaA1.2LP probe yielded a high-confidence set of 22 enriched proteins after subtracting no-probe controls across two probe concentrations. Beyond the cognate synthetase SpaKC-1, the top hits included two additional lanthipeptide synthetases, SpaKC-2 from a second class III BGC and SpaL from the class IV BGC, and two M1 family metallopeptidases, SpaP1 and SpaP2, whose encoding genes lie distally from any annotated RiPP BGC. In vitro assays confirmed that SpaP1 and SpaP2 cleave the leader peptide of cyclized SpaA1.2 at defined sites and process every lanthipeptide precursor encoded in S. sparsogenes, identifying them as promiscuous, extra-cluster processing proteases. Systematic cross-substrate testing further revealed graded cross-cluster activities: SpaKC-1 fully dehydrated the lan III-2 precursor SpaA2 but was inactive toward the class IV precursor SpaAL, while SpaKC-2 showed broader promiscuity toward all three lan III-1 precursors, and SpaL strongly preferred SpaAL with limited activity on SpaA1.2 and SpaA1.3. Coordinated temporal expression of all five genes during S. sparsogenes fermentation, each peaking on day five, supports the interpretation that these enzymes function within a coregulated, distributed modification network.
The combinatorial implications of this cross-cluster activity are considerable. Sequential processing by pairs of synthetases generates intermediates that are substrates for enzymes inactive toward the unmodified precursor. In one example, SpaKC-2 first dehydrates SpaA1.1 at Ser16 to produce a monocyclic intermediate that SpaL, which shows negligible activity toward the native precursor, then elaborates with six additional dehydrations. Across intra- and intercluster single- and multistep combinations, the three lanthipeptide BGCs in S. sparsogenes can collectively produce 17 structurally distinct products from a limited precursor set. The authors note that these activities have so far been demonstrated only in vitro or in heterologous hosts, and that targeted LC/MS analysis combined with synthetase deletion strains will be required to assess the in vivo relevance of cross-modified products. Looking ahead, the LP-probe framework is not confined to lanthipeptides: its successful application to the RiPP Recognition Element-containing cyanobactin enzyme LynD suggests extensibility to any RiPP pathway governed by leader-peptide recognition, and the authors propose that the approach will be especially valuable for organisms, including plants and animals, where dispersed gene architectures confound genome-based pathway annotation.