Antimicrobial resistance has emerged as one of the defining challenges in modern medicine, renewing scientific interest in antimicrobial peptides, AMPs, as alternatives to conventional antibiotics. These short molecules, found in the innate immune systems of nearly all living organisms, are best known for disrupting bacterial membranes. Yet a persistent puzzle has surrounded the wide gap between the peptide concentrations that perturb model lipid bilayers and the far higher amounts required to kill intact bacteria. This mismatch has long suggested that the bacterial cell envelope holds undiscovered molecular targets, and understanding those interactions could transform how researchers design next-generation peptide antibiotics.
Researchers supervised by Professor Isabelle Marcotte at the Université du Québec à Montréal, published in J. Am. Chem. Soc., investigated three amphibian-derived AMPs with distinct membrane-disruption mechanisms: aurein 1.2, caerin 1.1, and DMS-DA6. The team grew isotopically labeled cultures of Bacillus subtilis and Staphylococcus aureus, extracted purified cell walls, and probed AMP interactions using solid-state NMR, ssNMR, along with circular dichroism, turbidity assays, and solution-state 1H NMR. By combining 13C and 31P NMR experiments, the researchers could separately track rigid and flexible cell wall components at atomic resolution.
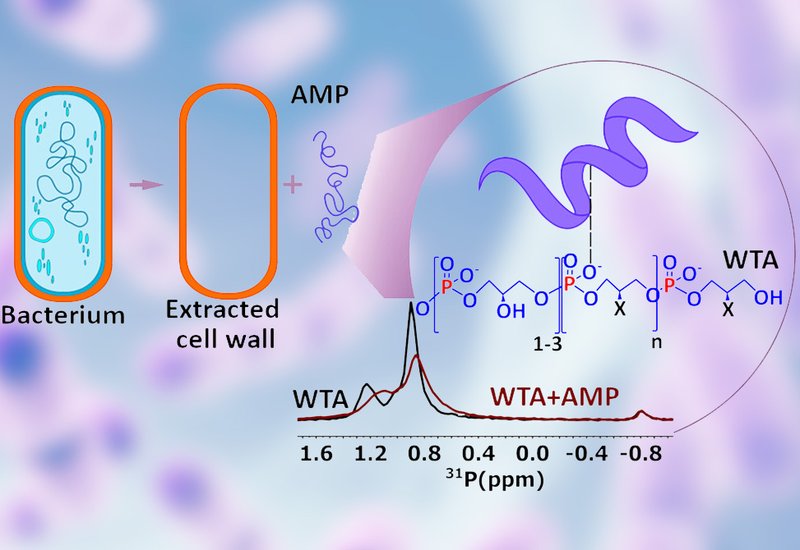
The 13C and 31P NMR data converged on a striking finding: all three AMPs bound preferentially to wall teichoic acids, WTAs, the anionic phosphate-rich polymers embedded in the Gram-positive cell wall, rather than to peptidoglycan. T2 relaxation measurements showed significant reductions in the mobility of WTA phosphate groups and glycan backbones upon AMP addition, consistent with aggregation and slowed collective motion. This dynamic change was visible at the mesoscopic scale as well, with optical microscopy revealing that aurein and caerin induced large-scale clumping of purified cell wall material while DA6 produced smaller aggregates. Solution NMR chemical shift mapping identified specific amino acid residues responsible for WTA recognition in each peptide, though no single conserved binding motif emerged across the three sequences. Circular dichroism experiments revealed that aurein adopted an alpha-helical conformation in the presence of WTAs, suggesting that cell wall binding triggers structural pre-organization before membrane engagement. Adding potassium chloride to block the charged WTA phosphates abolished this helical induction, confirming that electrostatic attraction drives the initial recognition event. DA6 showed strong WTA binding in both bacterial strains, while caerin, the most hydrophobic peptide of the three, interacted only weakly with isolated WTAs despite its potent antibacterial activity.
These results broaden the established model of AMP action beyond membrane permeabilization, casting WTAs as key early molecular targets in Gram-positive bacteria. The findings offer a mechanistic explanation for a well-known resistance strategy, as bacteria that increase the d-alanylation of WTAs reduce their net negative surface charge and thereby lower AMP affinity. The work also supports a multistep model in which AMPs first accumulate at the cell wall through electrostatic capture, in some cases undergoing structural changes, before gaining access to the cytoplasmic membrane for final disruption. This framework opens a rational path toward designing AMPs with tailored specificity for particular bacterial species or resistance profiles.